November 08, 2019
Hanasaki M. and Masumoto H. have developed a new tool to control gene induction in budding yeast, named as CRISPR/Transposon gene induction (CRITGI) at Biomedical Research Support Center (BRSC).
Hanasaki M. and Masumoto H. have developed a new tool to control gene induction in budding yeast, named as CRISPR/Transposon gene induction (CRITGI). CRITGI integrated an expression vector into Ty1 retrotransposon loci, and uses an amino acid to manage the gene transcription in a retrotransposon-dependent manner. This research has been published in Scientific Reports at 25th Oct 2019.
The fine-tuning of gene expression contributes to both basic science and applications. Here, we develop a novel gene expression technology termed CRITGI (CRISPR/Transposon gene integration). CRITGI uses CRISPR/Cas9 to integrate multiple copies of the plasmid pTy1 into Ty1 loci, budding yeast retrotransposons. The pTy1 plasmid harbors a Ty1 consensus sequence for integration, a gene of interest with its own promoter and a selection marker gene. Interestingly, the expression of the pTy1 gene in Ty1 loci could be induced in synthetic complete amino acid depletion medium, which could activate the selection marker gene on pTy1. The induction or repression of the gene on pTy1 depended on Ty1 transcription. Activation of the selection marker gene on pTy1 triggered Ty1 transcription, which led to induction of the gene on pTy1. The gene on pTy1 was not transcribed with Ty1 mRNA; the transcription required its own promoter. Furthermore, the trimethylation of histone H3 on lysine 4, a landmark of transcriptionally active chromatin, accumulated at the 5′ end of the gene on pTy1 following selection marker gene activation. Thus, CRITGI is a unique gene regulation system to induce the genes on pTy1 in amino acid depletion medium and utilizes Ty1 transcription to create a chromatin environment favorable for the transcription of the genes on pTy1.
Miki Hanasaki and Hiroshi Masumoto,
CRISPR/Transposon gene integration (CRITGI) can manage gene expression in a retrotransposon-dependent manner
Scientific Reports, 9, Article number: 15300 (2019)
URL: https://www.nature.com/articles/s41598-019-51891-6
|
|
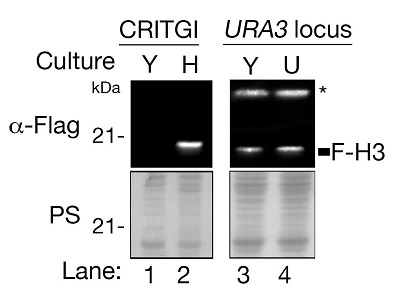
Y: Yeast-polypeptone-dextrose (YPD) containing histidine, H: SD-His (synthetic medium depleting histidine). CRITGI integrated Flag-HHT1 gene encoding Flag-histone H3 (F-H3) into Ty1 loci, and could express F-H3 in SD-His medium, although not in YPD medium. Flag-HHT1 gene integrated at URA3 locus was expressed in both SD-His and YPD media (control) |
CONTACT INFORMATION:
Hiroshi Masumoto, M.D.
Senior Assistant Professor, Biomedical Research Support Center (BRSC),
Nagasaki University School of Medicine
PHONE : +81-95-819-7089
E-MAIL : himasumo*nagasaki-u.ac.jp (change"*"to"@")